Nonparametric estimation of time-varying reproduction numbers
Daniel J. McDonald
Department of Statistics, The University of British Columbia

Mathematical modelling of disease / epidemics is very old
- Daniel Bernoulli (1760) - studies inoculation against smallpox
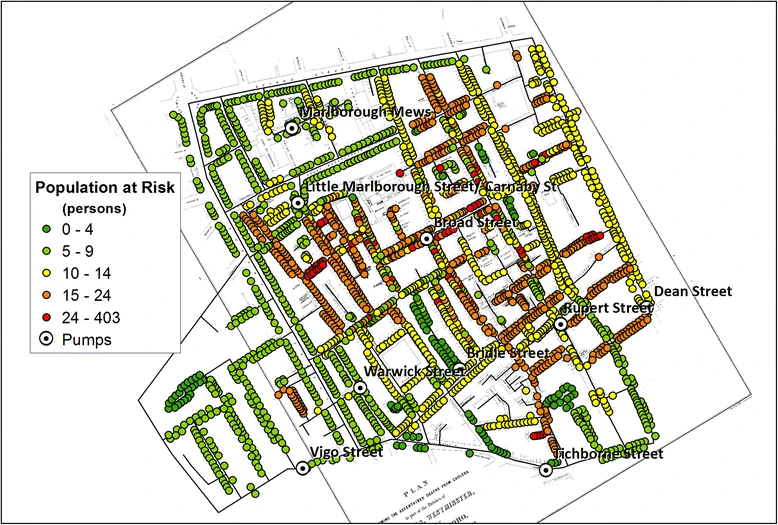
- John Snow (1855) - cholera epidemic in London tied to a water pump
- Ronald Ross (1902) - Nobel Prize in Medicine for work on malaria
- Kermack and McKendrick (1927-1933) - basic epidemic (mathematical) model
\(R_0\) basic reproduction number
Dates at least to Alfred Lotka (1920s) and others (Feller, Blackwell, etc.)
The expected number of secondary infections due to a primary infection
- \(R_0 < 1\) ⟹ the epidemic will die out
- \(R_0 > 1\) ⟹ the epidemic will grow until everyone is infected
\(R_0\) is entirely retrospective
It’s a property of the pathogen in a fully susceptible (infinite) population
Each outbreak is like a new sample
To estimate something like this from data, the “bonehead” way is to
- Wait until the epidemic is over (no more infections circulating)
- Contact trace the primary infection responsible for each secondary infection
- Take a sample average of the number caused by each primary
- Possibly repeat over many outbreaks
Of course no one actually does that
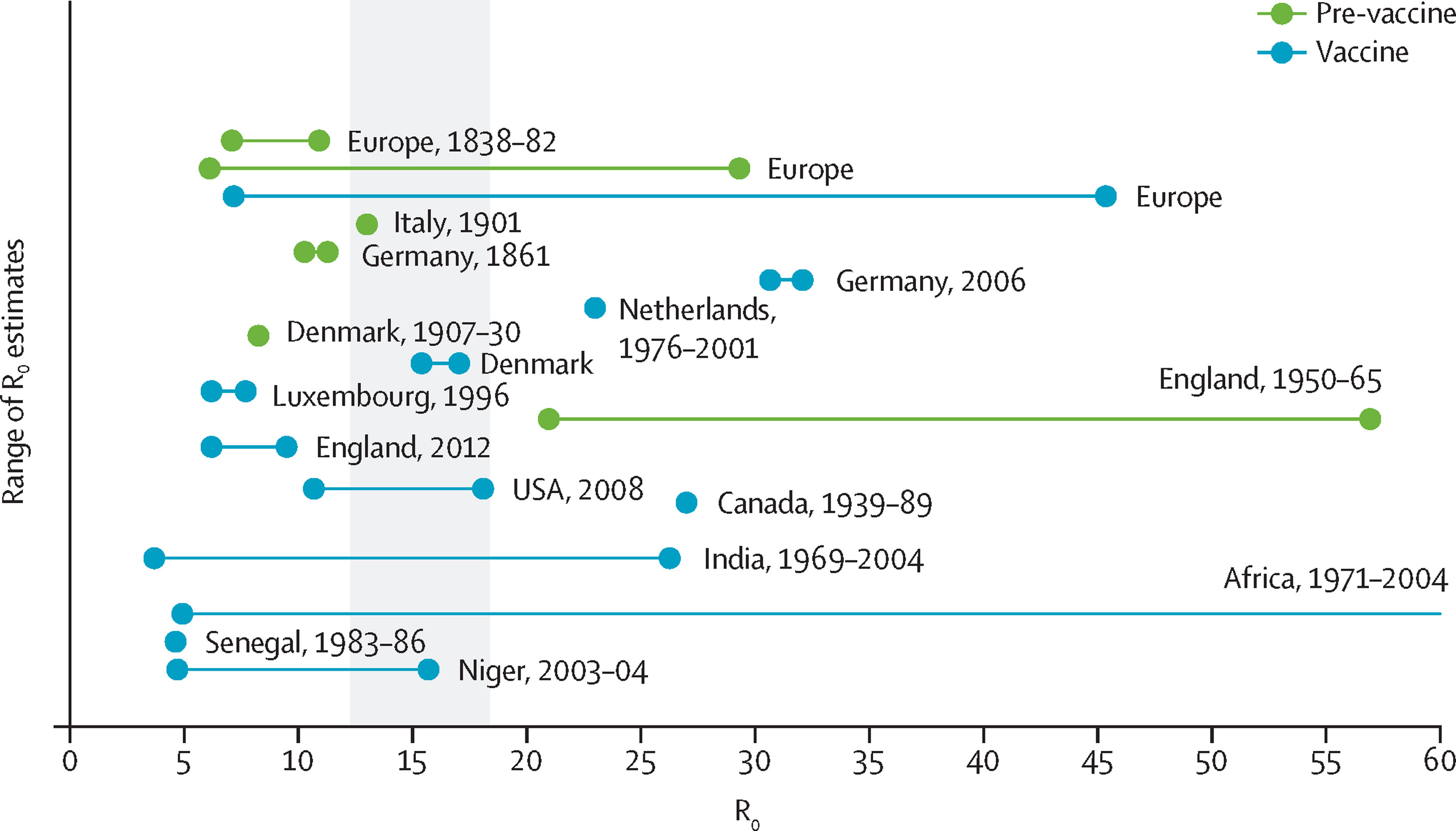
Lots of work on how to estimate \(R_0\)
Effective reproduction number
Suppose \(s\)% of the population is susceptible
Then, “the” effective reproduction number \(R=sR_0\)
Allows you to reason about things like
The level of vaccine coverage necessary to prevent an outbreak from growing uncontrollably.
So, for measles, if \(R_0\approx 15\), the disease will die out if immunity is
\[ sR_0\ \leq\ 1\ \Longrightarrow\ 1-s\ \leq\ 1-\frac{1}{R_0}\ \approx\ 93\% \]
\(R(t)\) — instantaneous reproduction number
- The effective reproduction number in the middle of an outbreak
- Some of the population is immune, others are infected, others susceptible
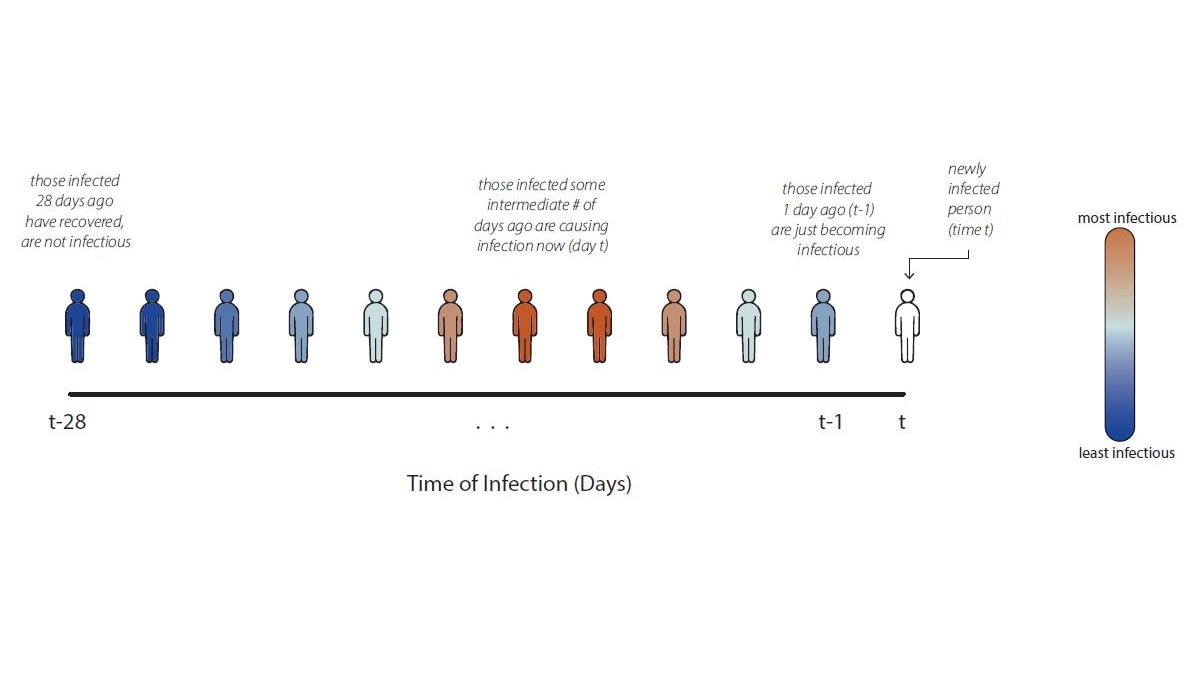
The expected number of secondary infections at time \(t\) caused by an earlier primary infection
\(f(a) \geq 0,\ \forall a \geq 0\) — the rate at which an infection of age \(a\) produces new infections
\(\displaystyle g(a) := \frac{f(a)} {\int_{0}^\infty f(a)\ \mathsf{d}a} = \frac{f(a)}{R_0}.\)
\(\displaystyle R_0 := \int_{0}^\infty f(a)\ \mathsf{d}a.\)
Can allow \(g(t, a)\), hold this fixed for now.
The generation interval distribution \(g(a)\)
SIR-type (compartmental) models - Stochastic Version
Suppose each of N people in a bucket at time t:
Susceptible(t) : not sick, but could get sick
Infected(t) : sick, can make others sick
Removed(t) : recovered or dead; not sick, can’t get sick
- During period \(h\), each \(S\) meets \(kh\) people.
- Assume \(\mathbb{P}( S \textrm{ meets } I \textrm{ and becomes } I ) = c\).
- Then \(\mathbb{P}( S(t) \rightarrow I(t+h) ) = 1 - (1 - c I(t) / N )^{hk} \approx kchI(t) / N\).
- Therefore, \(I(t+h) \mid S(t),\ I(t) \sim \textrm{Binom}(S(t),\ kchI(t) / N)\).
- Assume \(\mathbb{P}( I(t) \rightarrow R(t+h)) = \gamma h,\ \forall t\).
- Then \(R(t+h) \mid I(t) \sim \textrm{Binom}(I(t),\ \gamma h)\).
SIR-type (compartmental) models - Stochastic Version
\[\begin{aligned} x(t+h) &\sim \mathrm{Binom}\left(S(t),\ \frac{\beta}{N} h I(t)\right) = \text{incident infections}\\ z(t+h) &\sim \mathrm{Binom}\left(I(t),\ \gamma h\right) = \text{incident recoveries}\\ S(t+h) & = S(t) - x(t+h)\\ I(t+h) & = I(t) + x(t+h) - z(t+h)\\ R(t+h) & = R(t) + z(t+h) \end{aligned}\]
In the deterministic limit, \(h\rightarrow 0\)
\[\begin{aligned} \frac{\mathsf{d}}{\mathsf{d}t}S(t) & = -\frac{\beta}{N} S(t)I(t)\\ \frac{\mathsf{d}}{\mathsf{d}t}I(t) & = \frac{\beta}{N} I(t)S(t) - \gamma I(t)\\ \frac{\mathsf{d}}{\mathsf{d}t}R(t) & = \gamma I(t) \end{aligned}\]
THE SIR model is often ambiguous between these.
Typically, people mean the deterministic, continuous time version.
\(R_t\) in compartmental models
There is an equivalence between a compartmental model and the renewal equation.
\[\begin{aligned} R_0 &= \beta\ /\ \gamma\\ x_{t+1} &= \beta \frac{S_{t}}{N} \sum_{k = 0}^{t-1} \big[(1-\gamma)^{k}\big] x_{t-k} = R_0 \frac{S_{t}}{N} \sum_{k = 0}^{t-1} \big[\gamma(1-\gamma)^{k}\big] x_{t-k} = R_{t}\sum_{k = 0}^{t-1} g(k)x_{t-k} \end{aligned}\]
\(R(t)\) and \(R_t\) — renewal equation
Alternatively: rather than working with a particular compartmental model
Define \(R(t)\) implicitly through the renewal equation
\[ x(t) = R(t)\int_0^\infty x(t-a)g(a)\mathsf{d}a = R(t)\times (x * g)(t), \]
where \(x(t)\) are infections at time \(t\).
In discrete time,
\[ x_{t+1} = R_t\sum_{a=0}^\infty x_{t-a}\widetilde{g}_a = R_t\times (x * \widetilde{g})_t. \]
And stochasticly,
\[ \mathbb{E}\big[x_{t+1} \mid x_1,\ldots,x_{t}\big] = R_t\sum_{a=0}^\infty x_{t-a}\widetilde{g}_a = R_t\times (x * \widetilde{g})_t. \]
Most estimators start here: \[ \mathbb{E}\big[x_{t+1} \mid x_1,\ldots,x_{t}\big] = R_t\sum_{a=0}^\infty x_{t-a}\widetilde{g}_a. \]
- Assume \(\widetilde{g}\) is known
- Model \(x_{t+1} \mid x_1,\ldots,x_{t}\) as Poisson or Negative Binomial
- Turn some inferential crank
\(R_t\) for Influenza
Source: US CDC Center for Forecasting Analytics
Estimating \(R_t\)
Data issues
\(x_t\) is incident infections, but we don’t ever see those
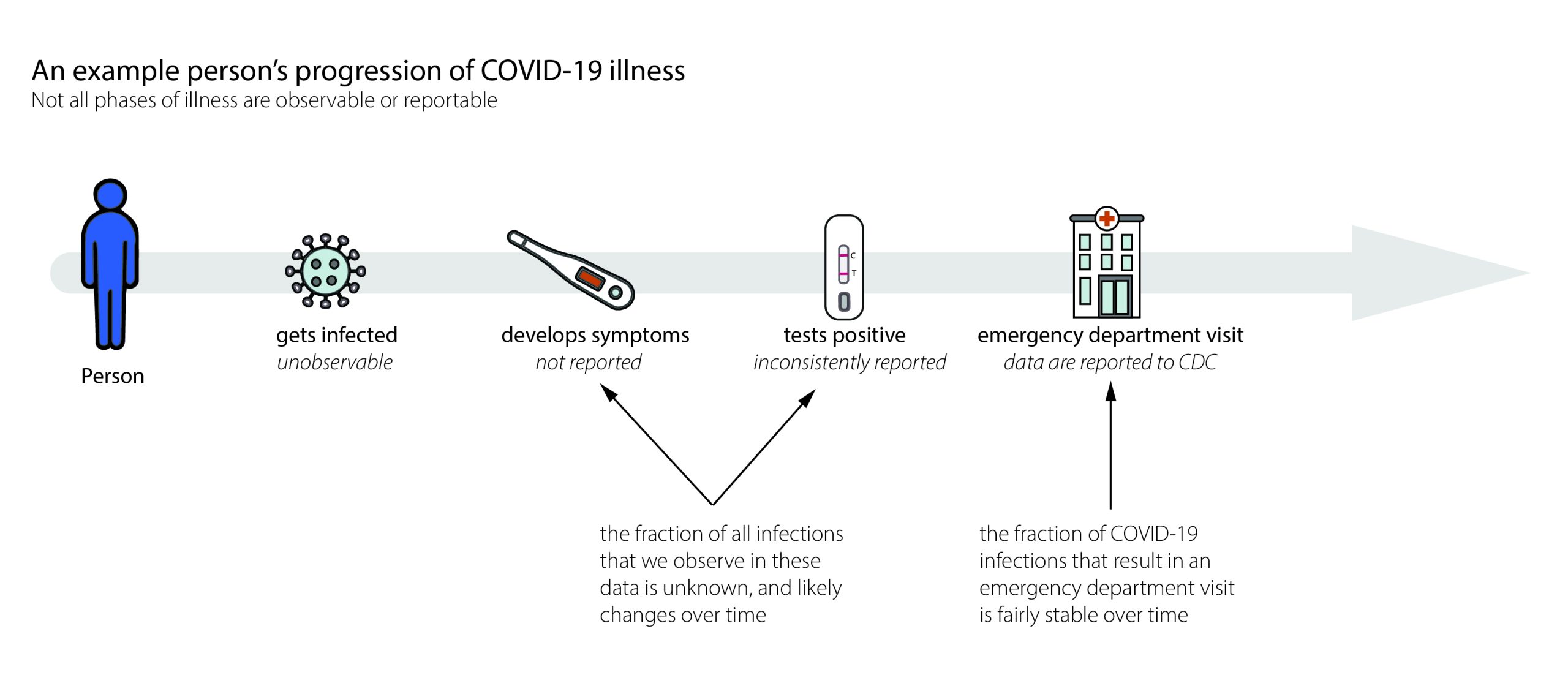
- Replace infections with cases
- Replace generation interval with serial interval
- Assume we have the serial interval
Serial interval distribution
Standard model for \(R_t\)
\[\begin{aligned} \eta_t &= \sum_{a=0}^{t-1} y_{t-a}p_a,\\ \\ y_t\ |\ y_{t-1},\ldots,y_{1} &\sim \textrm{Poisson}(R_t\eta_t). \end{aligned}\]
- Using \(y\) instead of \(x\) to be cases or hospitalizations or deaths, incidence
- Using \(p\) for serial interval distribution (discretized)
- The MLE for \(R_t\) is just \(y_t / \eta_t\).
- This has really high variance, but unbiased.
- So everybody smooths it.
This isn’t going to give you “\(R_t\)”, because you didn’t use incident infections.
At least, the timing is wrong, because the observable happened after infection.
Better to deconvolve incidence first (LSM2026).
The state of the art
{EpiEstim}CFFC2013
- Gamma prior on \(R_t\), but use a trailing window
- Super fast computationally
- Smoothness is ad hoc
{EpiFilter}Parag 2020
- State space model
- One step smoothness: \(R_{s+1} \sim \textrm{Gaussian}(R_s,\ \alpha R_s)\)
- Uses a discretized particle filter-type algorithm
{EpiLPS}GFH2023
- Negative Binomial likelihood
- Smoothness via \(\log(R_t\eta_t) = \mathbf{B}_{t,:}\beta\)
- \(\mathbf{B}\) is cubic B-spline basis, weighted Ridge penalty on \(\beta\)
- More priors, use Metropolis Adjusted Langevin Algorithm
{EpiNow2} CDC + CFA, Abbott, et al., 2023
- Negative Binomial likelihood
- Smoothness via a GP prior
- Accommodates the sequence of delays from infection \(\longrightarrow\) ??
- Adjusts for real-time issues like partial reporting
- Big Bayesian MCMC in Stan. Very slow.
Our model
Let \(\theta_t := \log(R_t)\).
Use Poisson likelihood.
\[ \begin{aligned} \widehat{R} &= \exp(\widehat{\theta}) &\widehat{\theta} &= \mathop{\mathrm{argmin}}_\theta\; \eta^{\mathsf{T}}\exp(\theta) - \mathbf{y}^{\mathsf{T}}\theta + \lambda\Vert D^{(k+1)}\theta\Vert_1 \end{aligned} \]
- Convex, has a global optimum
- \(\lambda\) controls smoothness relative to data fidelity
- \(\ell_1\) penalty produces adaptive piecewise polynomials of order \(k+1\)
- Near minimax optimal for functions with bounded total variation
Local adaptivity — \(\ell_1\) vs. \(\ell_2\)
Polynomial order, \(k=0\)
\[
\begin{aligned}
D^{(1)} &= \begin{bmatrix}
1 & -1 & & & & \\
& 1 & -1 & & & \\
& & & \ddots && \\
& & & & 1 & -1
\end{bmatrix} \\ \\
&\in \mathbb{R}^{(n-1)\times n}
\end{aligned}
\]
Polynomial order, \(k=1\)
\[
\begin{aligned}
D^{(2)} &= \begin{bmatrix}
1 & -2 & 1 & & & \\
& 1 & -2 & 1 & & \\
& & & \ddots && \\
& & & 1 & -2 & 1
\end{bmatrix} \\ \\
&= D^{(1)}D^{(1)}\\ \\
&\in \mathbb{R}^{(n-k-1)\times n}
\end{aligned}
\]
Polynomial order, \(k=2\)
\[\begin{aligned}
D^{(3)} &= \begin{bmatrix}
-1 & 3 & -3 & 1 & & \\
& -1 & 3 & -3 &1 & \\
& & & \ddots && \\
& & -1 & 3 & -3 & 1
\end{bmatrix} \\ \\
&= D^{(1)}D^{(2)}\\ \\
&\in \mathbb{R}^{(n-k-1)\times n}
\end{aligned}\]
Estimation algorithm
\[\mathop{\mathrm{minimize}}_\theta\; \eta^{\mathsf{T}}\exp(\theta) - \mathbf{y}^{\mathsf{T}}\theta + \lambda\Vert D^{(k+1)}\theta\Vert_1\]
Estimation algorithm
\[\mathop{\mathrm{minimize}}_{\theta,\ {\color{BurntOrange} \alpha}}\; \eta^{\mathsf{T}}\exp(\theta) - \mathbf{y}^{\mathsf{T}}\theta + \lambda\Vert D^{(1)}{\color{BurntOrange} \alpha}\Vert_1\quad {\color{BurntOrange} \textrm{subject to}\quad \alpha = D^{(k)}\theta}\]
Estimation algorithm
\[\mathop{\mathrm{minimize}}_{\theta,\ \alpha}\; \eta^{\mathsf{T}}\exp(\theta) - \mathbf{y}^{\mathsf{T}}\theta + \lambda\Vert D^{(1)} \alpha\Vert_1\quad \textrm{subject to}\quad \alpha = D^{(k)}\theta\]
Alternating direction method of multipliers (ADMM)
\[\begin{aligned} \theta &\longleftarrow \mathop{\mathrm{argmin}}_\theta\ \eta^{\mathsf{T}}\exp(\theta) - \mathbf{y}^{\mathsf{T}}\theta + \frac{\rho}{2}\Vert D^{(k)}\theta - \alpha + u \Vert_2^2 \\ \alpha &\longleftarrow \mathop{\mathrm{argmin}}_\alpha\ \lambda\Vert D^{(1)} \alpha \Vert_1 + \frac{\rho}{2}\Vert D^{(k)}\theta - \alpha + u \Vert_2^2 \\ u &\longleftarrow u + D^{(k)}\theta - \alpha \end{aligned}\]
Estimation algorithm
\[\mathop{\mathrm{minimize}}_{\theta,\ \alpha}\; \eta^{\mathsf{T}}\exp(\theta) - \mathbf{y}^{\mathsf{T}}\theta + \lambda\Vert D^{(1)} \alpha\Vert_1\quad \textrm{subject to}\quad \alpha = D^{(k)}\theta\]
Alternating direction method of multipliers (ADMM)
\[\begin{aligned} \theta &\longleftarrow \Big[{\color{Cerulean}\textrm{Proximal Newton / Fisher Scoring}}\Big] \\ \alpha &\longleftarrow \Big[{\color{BurntOrange}\textrm{Fused Lasso Signal Approximator}}\Big] \\ u &\longleftarrow u + D^{(k)}\theta - \alpha \end{aligned}\]
Solve sequentially for \(\Vert (D^{\dagger})^{\mathsf{T}}(\eta - y)\Vert_\infty = \lambda_1 > \cdots > \lambda_M=\epsilon \lambda_1\).
Results and features
Canadian Covid-19 cases
\(R_t\) for Canadian Covid-19 cases
Reconvolved Canadian Covid-19 cases
Example simulations for different methods
{rtestim} software
Guts are in C++ for speed
Lots of the usual S3 methods
Approximate “confidence” bands
\(\widehat{R}\) is a member of a function space
Arbitrary spacing of observations
Built-in cross validation
Time-varying delay distributions
{rtestim} software
Guts are in C++ for speed
Lots of the usual S3 methods
Approximate “confidence” bands
\(\widehat{R}\) is a member of a function space
Arbitrary spacing of observations
Built-in cross validation
Time-varying delay distributions
Approximation + Delta method gives
\[ \textrm{Var}(\widehat{R}) = \left(\textrm{diag}(\widehat{y}) + \lambda D^{\mathsf{T}}D\right)^{\dagger} \left(\frac{1}{\eta^2}\right) \]
{rtestim} software
Guts are in C++ for speed
Lots of the usual S3 methods
Approximate “confidence” bands
\(\widehat{R}\) is a member of a function space
Arbitrary spacing of observations
Built-in cross validation
Time-varying delay distributions
The solution is an element of the space of discrete splines of order \(k\) Tibshirani 2020
- Lets us interpolate (and extrapolate) to off-observation points
- Lets us handle uneven spacing
{rtestim} software
Guts are in C++ for speed
Lots of the usual S3 methods
Approximate “confidence” bands
\(\widehat{R}\) is a member of a function space
Arbitrary spacing of observations
Built-in cross validation
Time-varying delay distributions
{rtestim} software
Summary and collaborators
Contributions
- Framework for \(R_t\) estimation
- Naturally imposes smoothness
- Fast, extensible, easy to use
- Has “confidence” bands
Future work
- Allow Negative Binomial likelihood
- Deconvolve incidence
- Forecast
- Mixed smoothness behaviours
- Figure out CV for convolution
- Better confidence bands
- Use relationship with growth rate
Collaborators and funding





{rtestim} — dajmcdon/rtestim-2025